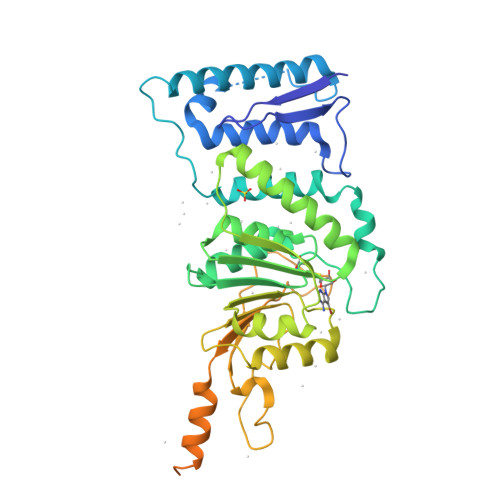
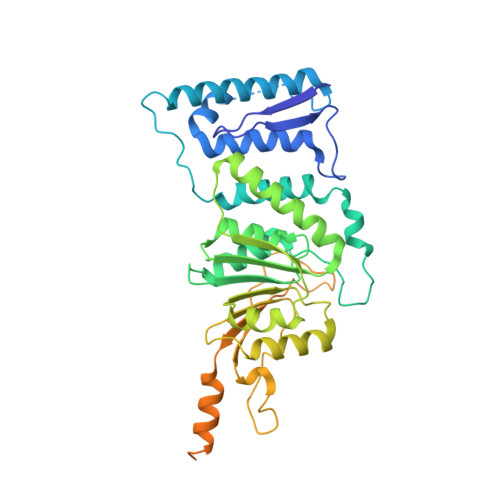
Bromo-deaza-SAH: a potent and selective DOT1L inhibitor.
Yu, W., Smil, D., Li, F., Tempel, W., Fedorov, O., Nguyen, K.T., Bolshan, Y., Al-Awar, R., Knapp, S., Arrowsmith, C.H., Vedadi, M., Brown, P.J., Schapira, M.(2013) Bioorg Med Chem 21: 1787-1794
- PubMed: 23433670
- DOI: https://doi.org/10.1016/j.bmc.2013.01.049
- Primary Citation of Related Structures:
3SX0 - PubMed Abstract:
Chemical inhibition of proteins involved in chromatin-mediated signaling is an emerging strategy to control chromatin compaction with the aim to reprogram expression networks to alter disease states. Protein methyltransferases constitute one of the protein families that participate in epigenetic control of gene expression, and represent a novel therapeutic target class. Recruitment of the protein lysine methyltransferase DOT1L at aberrant loci is a frequent mechanism driving acute lymphoid and myeloid leukemias, particularly in infants, and pharmacological inhibition of DOT1L extends survival in a mouse model of mixed lineage leukemia. A better understanding of the structural chemistry of DOT1L inhibition would accelerate the development of improved compounds. Here, we report that the addition of a single halogen atom at a critical position in the cofactor product S-adenosylhomocysteine (SAH, an inhibitor of SAM-dependent methyltransferases) results in an 8-fold increase in potency against DOT1L, and reduced activities against other protein and non-protein methyltransferases. We solved the crystal structure of DOT1L in complex with Bromo-deaza-SAH and rationalized the observed effects. This discovery reveals a simple strategy to engineer selectivity and potency towards DOT1L into the adenosine scaffold of the cofactor shared by all methyltransferases, and can be exploited towards the development of clinical candidates against mixed lineage leukemia.
Organizational Affiliation:
Structural Genomics Consortium, University of Toronto; 101 College Street, MaRS Centre, South tower, Toronto, Ontario, M5G 1L7, Canada.